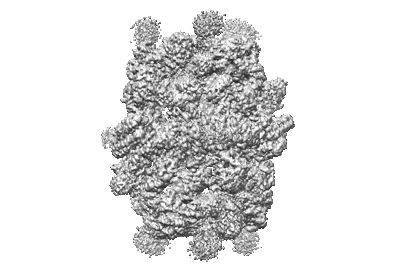
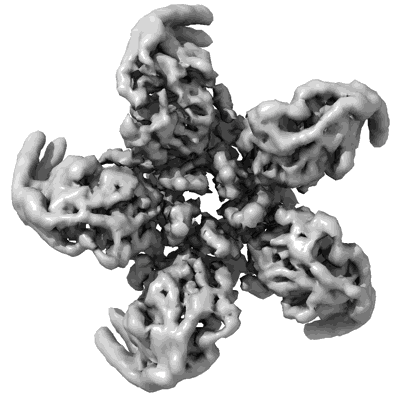
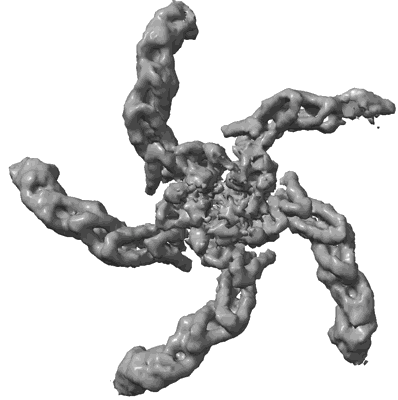
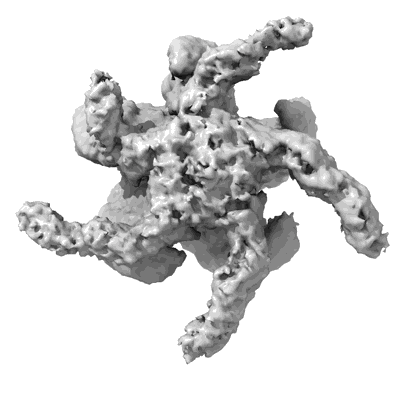
CryoEM structure of the HisRS-like domain of human GCN2
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8t7t
Q-score: 0.45
Yin JZ, Keszei AFA, Mazhab-Jafari MT, Sicheri F
To Be Published
- Dimeric hisrs-like domain of human gcn2 (Complex from Homo sapiens)
- Eif-2-alpha kinase gcn2 (187 kDa, Protein from Homo sapiens)
MBP-tagged fusion fatty acid synthase from S. cerevisiae
Samani EK, Chen AC, Lou JW, Dai DL, Keszei AFA, Tan G, Boone C, Grininger M, Mazhab-Jafari MT
Commun Biol (2024) 7 pp. 92- [ DOI: doi:10.1038/s42003-024-05777-7 ]
MBP tagged and Mixed chain fatty acid synthase, FAS2_deltaACP
Samani EK, Chen AC, Lou JW, Dai DL, Keszei AFA, Tan G, Boone C, Grininger M, Mazhab-Jafari MT
Commun Biol (2024) 7 pp. 92- [ DOI: doi:10.1038/s42003-024-05777-7 ]
MBP tagged mixed chain FAS, fusFAS_deltaACP
Samani EK, Chen AC, Lou JW, Dai DL, Keszei AFA, Tan G, Boone C, Grininger M, Mazhab-Jafari MT
Commun Biol (2024) 7 pp. 92- [ DOI: doi:10.1038/s42003-024-05777-7 ]
Structure and dynamics of a pentameric KCTD5/Cullin3/GBeta1Gamma2 E3 ubiquitin ligase complex
Nguyen DM, Rath DH, Devost D, Pétrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hébert TE, Privé GG
Proc Natl Acad Sci U S A (2024) 121 [ DOI: 10.1073/pnas.2315018121 Pubmed: 38625940 ]
Image sets:
- K5C3Gbg1 Krios Data Grid 2 multiframe micrographs
- K5C3Gbg Particles
- K5C3Gbg Krios Data Grid 2 TILT multiframe micrographs. Collected with 40 degree tilt.
- MotionCorrected Data for K5C3Gbg Grid2 TILT
- K5C3Gbg1 Krios Data Grid 1 multiframe micrographs
- MotionCorrected Data for normal(Grid1+Grid2) K5C3Gbg
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(CTD)/Gbeta1gamma2
Nguyen DM, Rath DH, Devost D, Petrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hebert TE, Prive GG
PNAS (2024) 121 pp. e2315018121-e2315018121 [ DOI: doi:10.1073/pnas.2315018121 Pubmed: 38625940 ]
- Gbb1_human (Protein from Homo sapiens)
- Complex of substrate adaptor kctd5 with cullin-3 and substrate gbeta-gamma (Complex from Homo sapiens)
- Gbg2_human (Protein from Homo sapiens)
- Kctd5_human (Protein from Homo sapiens)
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(BTB)/Cullin3(NTD)
Nguyen DM, Rath DH, Devost D, Petrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hebert TE, Prive GG
PNAS (2024) 121 pp. e2315018121-e2315018121 [ DOI: doi:10.1073/pnas.2315018121 Pubmed: 38625940 ]
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Composite Map from RELION Multi-body Refinement
Resolution: 3.82 Å
EM Method: Single-particle
Fitted PDBs: 8u81
Q-score: 0.228
Nguyen DM, Rath DH, Devost D, Petrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hebert TE, Prive GG
PNAS (2024) 121 pp. e2315018121-e2315018121 [ DOI: doi:10.1073/pnas.2315018121 Pubmed: 38625940 ]
- Cullin3 1-381 (Protein from Homo sapiens)
- Gbg2_human (Protein from Homo sapiens)
- Gbb1_human (Protein from Homo sapiens)
- Kctd5_human (Protein from Homo sapiens)
- Complex of substrate adaptor kctd5 with cullin-3 and substrate gbeta-gamma (Complex from Homo sapiens)
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Reference Map for Composite Map EMD-41996
Nguyen DM, Rath DH, Devost D, Petrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hebert TE, Prive GG
PNAS (2024) 121 pp. e2315018121-e2315018121 [ DOI: doi:10.1073/pnas.2315018121 Pubmed: 38625940 ]
- Kctd5 (Protein from Homo sapiens)
- Cullin3 1-381 (Protein from Homo sapiens)
- Gbg2_human (Protein from Homo sapiens)
- Gbb1_human (Protein from Homo sapiens)
- Complex of substrate adaptor kctd5 with cullin-3 and substrate gbeta-gamma (Complex from Homo sapiens)
Nguyen DM, Rath DH, Devost D, Petrin D, Rizk R, Ji AX, Narayanan N, Yong D, Zhai A, Kuntz DA, Mian MUQ, Pomroy NC, Keszei AFA, Benlekbir S, Mazhab-Jafari MT, Rubinstein JL, Hebert TE, Prive GG
PNAS (2024) 121 pp. e2315018121-e2315018121 [ DOI: doi:10.1073/pnas.2315018121 Pubmed: 38625940 ]